In late November last year, epidemiologists began noticing some unusual patterns in the spread of COVID-19 in the South East of England.
A national lockdown had been in place since the start of the month, yet despite initial drop offs, cases in parts of Kent, Essex and London had started to increase again.
The problem was flagged to a small team of PHE scientists and external academics in the specialised field of viral genomics – the study of a virus’s genetic make-up.
They began looking at cases that had undergone sequencing (genetic analysis) across Kent. What they saw was lots of cases of a new variant that featured high numbers of unusual mutations.
Unbeknownst to the team at the time, this variant would set off on a rapid expansion across the South East of England, eventually taking over as the dominant COVID-19 strain in the UK and spreading to over 100 countries.
The investigation begins
Doctor Meera Chand, one of the national COVID-19 incident directors at PHE, and Richard Myers, Head of Bioinformatics at PHE, were among the first to become aware of the new variant, then known as B.1.1.7, and were part of the team tasked with quickly understanding what the UK was dealing with.
“We immediately knew we had found something very concerning”, says Meera. “Normally when you’re looking at samples you would expect to see lots of small clusters made up of multiple strains that are all slightly different. But when we looked at Kent, we saw about 50% of the samples were extremely similar, forming one massive cluster.”
Richard adds: “It was quite unusual. Up to that point, the virus had been characterised by having a relatively slow mutational rate, maybe two variations a month. Suddenly out of nowhere we’ve got this large cluster featuring mutations that were significantly different to what we’d looked at before.”
A team of experts from a range of disciplines across PHE and academic partners, including virology, epidemiology, modelling and genomics, was quickly assembled to study the characteristics of the Kent samples.
According to Meera: “It was really the first time we had brought together all the different disciplines needed to evaluate a variant at pace and understand how it behaves.”
The investigation immediately got underway, but there was a problem. The team knew which variant they were looking for, but data was limited to samples that had undergone whole genome sequencing (laboratory analysis that identifies a virus’s genetic make-up) – typically at the time around 5% of samples collected nationally.
A big breakthrough
Fortunately, the team were assisted by an unexpected source. Staff at ‘lighthouse labs’ (localised centres that test tens of thousands of swabs per day through an automated system) began noticing a problem with their test results coming from the Kent area.
One of the mutations in B.1.1.7 causes the ‘S-gene’, one of the characteristics identified when analysing the virus’s genetic code, to ‘drop out’, or in other words go missing.
Meera explains: “The lighthouse labs were having problems detecting the S-gene in their samples. Initially they thought there was an issue with the test. It wasn’t working, and the rate at which it wasn’t working was increasing very steeply.”
The team quickly began looking at the numbers of samples being tested across the lighthouse labs where the S-gene was missing. This widened the scope of their lens considerably, enabling them to identify cases of the new variant as they emerged.
“Identifying cases through the lighthouse labs was a huge leap ahead”, explains Meera. “Whole genome sequencing takes time, but lighthouse results are a real time way of detecting what’s happening. A huge proportion of samples go through them, so we were able to give good, real time monitoring of big data.
“It was suddenly very clear that the rate of spread was very rapid and was much bigger than just Kent. It was in London, Essex and other parts of the South East and expanding across the rest of the country.”
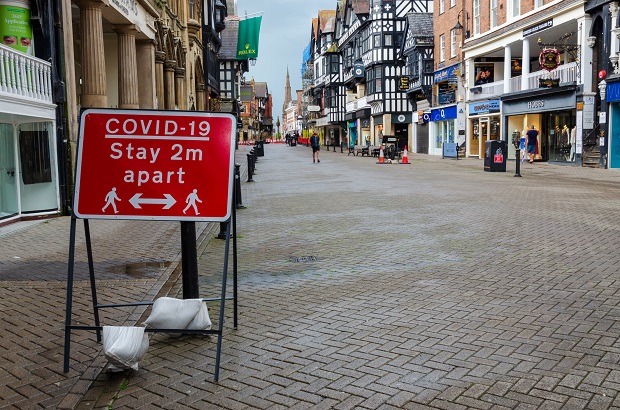

Acting fast to stop the spread
By December 21, the South East of England was again under tight COVID-19 restrictions, with much of the region placed in Tier 4. However, the rapid spread of B.1.1.7 meant hospitalisations and deaths were inevitable, putting the team under increasing pressure to quickly learn more about the threats the variant posed.
Richard explains how it felt: “It was obviously scientifically interesting to observe the mutations we were seeing, but we mustn’t ever forget the consequences of what we’re looking at. We were massively concerned to try and understand what the biological significance of these changes to the virus would be. There’s a sense of frustration in that work. It’s relatively straightforward to identify the changes, it’s a far more labour-intensive job to be able to say what they mean.”
Meera adds: “In mid-December, when all the analyses started coming together, we were really very worried. We realised what a bad situation we were potentially in and it was daunting to think about how much was resting on an analysis that was being done for the first time. I wouldn’t want to underestimate how challenging that time was for the team. However, this is why we had invested so much in genomics, to ensure we were well equipped to deal with this very scenario.”
Thanks to their early work, and the analysis of samples gathered in the lighthouse labs, the team were quickly able to establish to high degrees of confidence just how much more contagious the new variant was – now understood to be around 50% more than the original COVID-19 strain. This was able to be factored into public health decision making in December.
Further discoveries
Understanding the severity of B.1.1.7 (now referred to as VOC-20DEC-01) and its impact on hospitalisations and deaths however has been a more complex task. That picture continues to become clearer through the work of many independent academic groups across the country and, increasingly, internationally.
In January, a report by the New and Emerging Respiratory Virus Threats Advisory Group (NERVTAG) concluded a “realistic possibility” that infection with B.1.1.7 is associated with an increased risk of death, potentially by up to 30 per cent.
And a PHE report published in February on the effectiveness of vaccines found the Oxford/AstraZeneca and Pfizer jabs both provide high levels of immunity against the variant.
“B.1.1.7 is now our virus until something displaces it. Understanding that severity and, more recently, vaccine effectiveness was needed in order to build it into our planning for healthcare and treatments,” says Meera.
But despite major progress in understanding the behaviour of B.1.1.7, questions remain about the origin of the variant, and how and why such a range of mutations developed so quickly.
Richard explains: “I still don’t think we fully understand how and where that happened. A lot of the variants we observe in this country tend to be importation related, but in this one all evidence suggests this is probably a virus that has turned over in an individual in the UK for a long period of time and accumulated these mutations. It’s then popped out and started spreading, and by the time we saw it there were a significant number of cases.”
Looking ahead
Since the emergence of B.1.1.7, several other significant variants have been detected in the UK and are being tracked by PHE, though numbers remain low.
Meera says: “We are going to see more variants. The conditions are there globally, and we will need to monitor and assess them even more quickly than we are now. The most concerning characteristic right now would be vaccine escape, though in the medium term if it is needed, vaccine updates could be used as they are in the global system for flu.”
Domestically, genomic surveillance is expected to continue playing a leading role in informing national efforts to move out of restrictions by June 21, but international collaboration will also be essential for the future of the public health.
That global collaboration is already underway with the founding of the New Variant Assessment Platform, which aims to support the ability of countries around the world to identify variants of concern.
The platform is led by PHE, working with NHS Test and Trace, academic partners and the World Health Organization’s SARS-CoV-2 Global Laboratory Working Group.
And feeding into these wider programmes will be data and analysis gathered by all the individuals working in the ever-evolving fields of genomics and bioinformatics.
Richard says: “The scale of this pandemic has blown previous outbreaks like Swine Flu out of the water. The fact this has coincided with a certain level of maturity in terms of genomic sequencing and bioinformatic technology has had a huge impact on our ability to gather valuable data and learn about this virus.”
Meera adds: “The generation of viral genomic data in the UK has been an amazing development in the past year. And whilst we have to accept a bit of uncertainty in assessing a live situation, we need to use that data to provide the best insights we can in real time to help shape our response to the virus in this country going forward.”
Interested in reading more? Our blog on protecting ourselves as we move out of lockdown can be found here. You can subscribe to receive updates on new posts here.
View original article
Contributor: Richard Myers